Data visualization with ggplot2
Learning Objectives
- Produce scatter plots, boxplots, and time series plots using ggplot.
- Describe what faceting is and apply faceting in ggplot.
- Set universal plot settings.
We start by loading the required packages.
ggplot2 is included in the
tidyverse package.
library(tidyverse)Let’s read in our surveys data, but filter it to only get back rows
where ALL the data are present, also known as “complete cases”. We’re
also showing you a new little trick: using a period with a pipe.
Normally, a pipe just sends the stuff on the left into the FIRST
argument position in the function on the right. However, sometimes we
want that stuff to get sent to a slightly different place in the
righthand function. In this case, we want to send it into the
complete.cases() function, so that function will run on the
whole dataset. In order to specifically tell the pipe to send the
lefthand side into this function, we put a period there. You can think
of this as the target for the pipe.
surveys_complete <- read_csv("data/portal_data_joined.csv") %>%
filter(complete.cases(.))Plotting with ggplot2
ggplot2 is a plotting package that
makes it simple to create complex plots from data in a data frame. It
provides a more programmatic interface for specifying what variables to
plot, how they are displayed, and general visual properties. Therefore,
we only need minimal changes if the underlying data change or if we
decide to change from a bar plot to a scatterplot. This helps in
creating publication quality plots with minimal amounts of adjustments
and tweaking.
ggplot2 functions like data in the
‘long’ format, i.e., a column for every dimension, and a row for every
observation. Well-structured data will save you lots of time when making
figures with ggplot2
ggplot graphics are built step by step by adding new elements. Adding layers in this fashion allows for extensive flexibility and customization of plots.
To build a ggplot, we will use the following basic template that can be used for different types of plots:
ggplot(data = <DATA>, mapping = aes(<MAPPINGS>)) + <GEOM_FUNCTION>()- use the
ggplot()function and bind the plot to a specific data frame using thedataargument
ggplot(data = surveys_complete)- define a mapping (using the aesthetic (
aes) function), by selecting the variables to be plotted and specifying how to present them in the graph, e.g. as x/y positions or characteristics such as size, shape, color, etc.
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length))add ‘geoms’ – graphical representations of the data in the plot (points, lines, bars).
ggplot2offers many different geoms; we will use some common ones today, including:* `geom_point()` for scatter plots, dot plots, etc. * `geom_boxplot()` for, well, boxplots! * `geom_line()` for trend lines, time series, etc.
To add a geom to the plot use the + operator. Because we
have two continuous variables, let’s use geom_point()
first:
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point()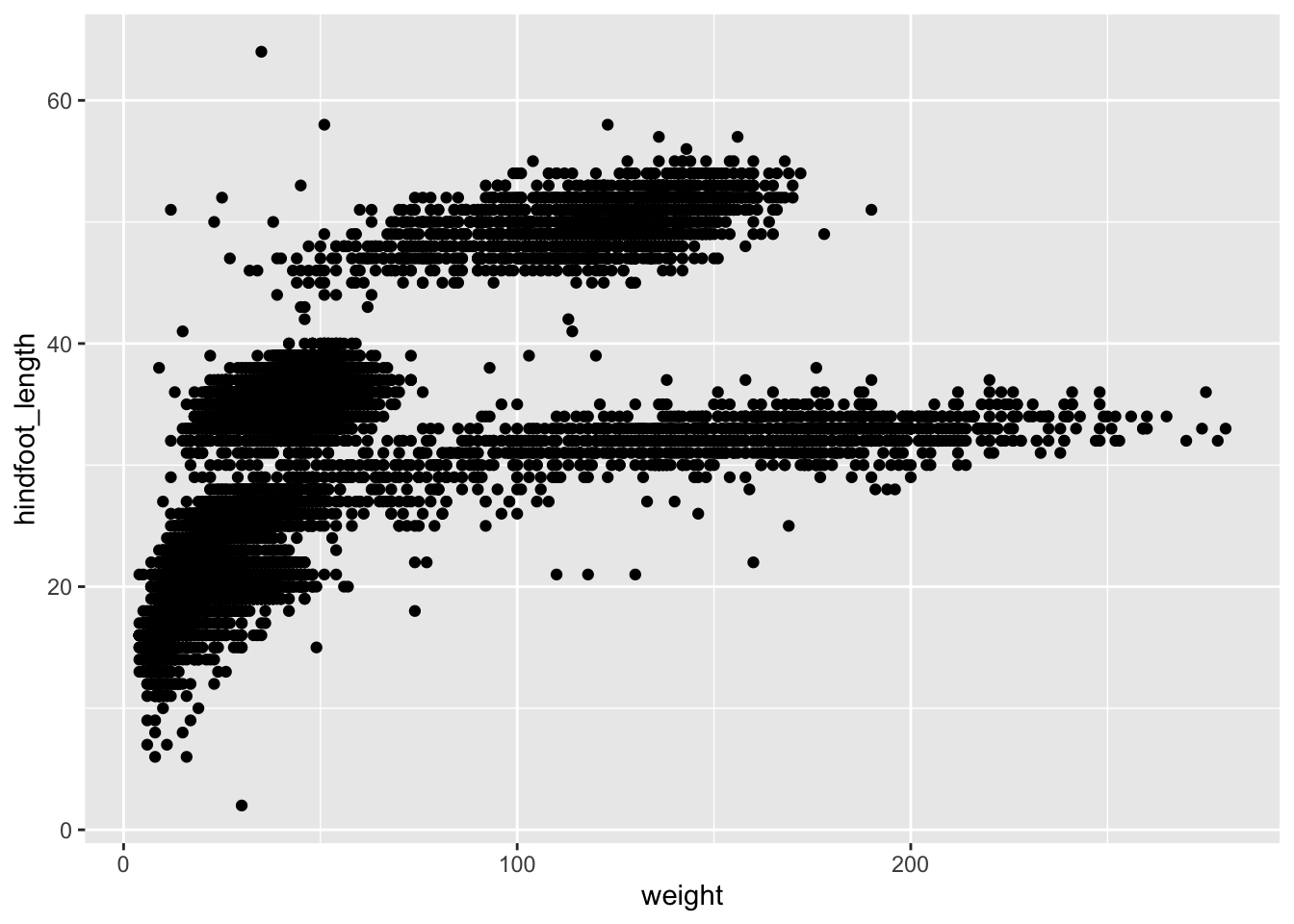
The + in the ggplot2
package is particularly useful because it allows you to modify existing
ggplot objects. This means you can easily set up plot
templates and conveniently explore different types of plots, so the
above plot can also be generated with code like this:
# Assign plot to a variable
surveys_plot <- ggplot(data = surveys_complete,
mapping = aes(x = weight, y = hindfoot_length))
# Draw the plot
surveys_plot +
geom_point()Notes
- Anything you put in the
ggplot()function can be seen by any geom layers that you add (i.e., these are universal plot settings). This includes the x- and y-axis mapping you set up inaes(). - You can also specify mappings for a given geom independently of the
mappings defined globally in the
ggplot()function. - The
+sign used to add new layers must be placed at the end of the line containing the previous layer. If, instead, the+sign is added at the beginning of the line containing the new layer,ggplot2will not add the new layer and will return an error message.
# This is the correct syntax for adding layers
surveys_plot +
geom_point()
# This will not add the new layer and will return an error message
surveys_plot
+ geom_point()Building your plots iteratively
Building plots with ggplot2 is
typically an iterative process. We start by defining the dataset we’ll
use, lay out the axes, and choose a geom:
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point()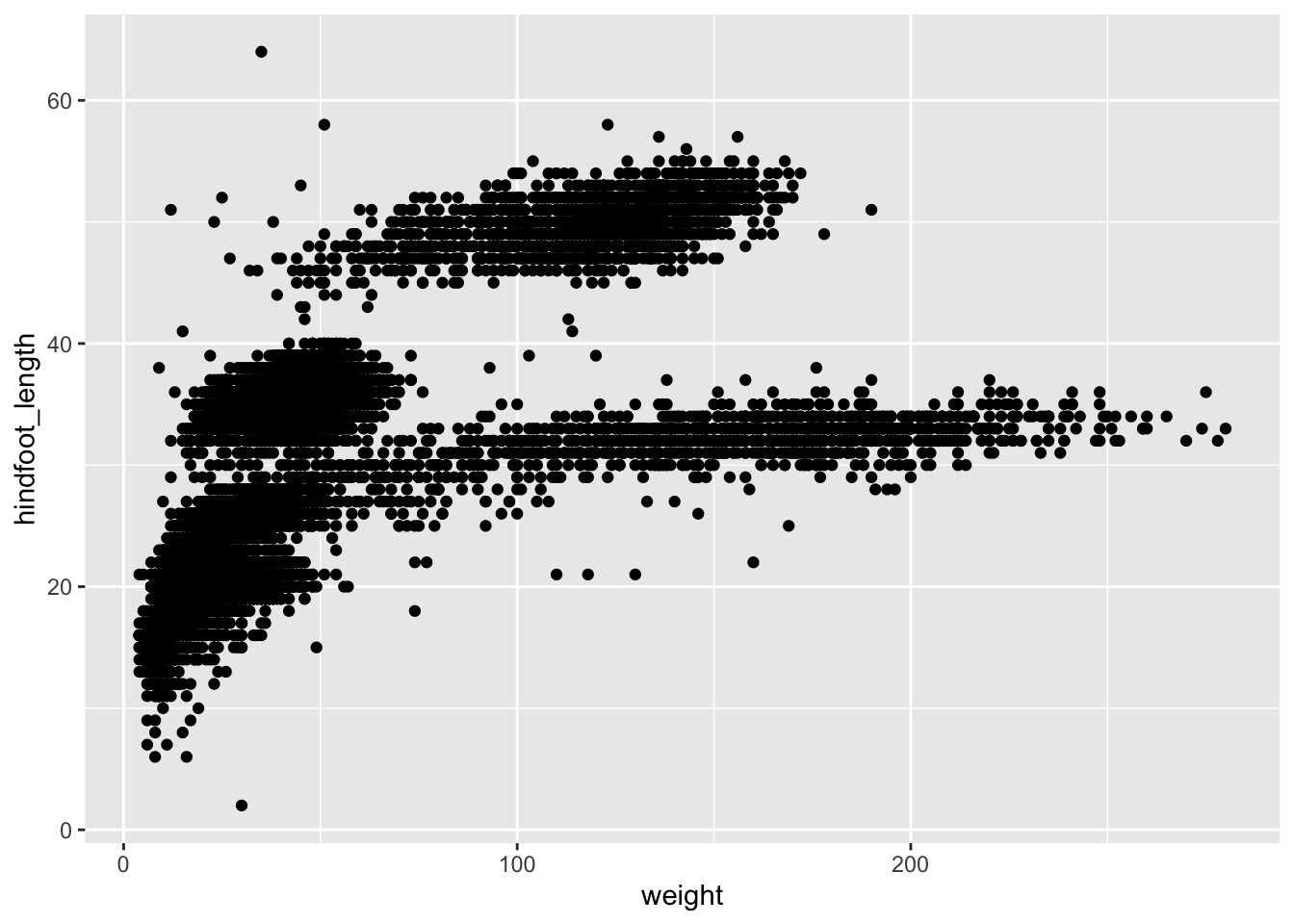
Then, we start modifying this plot to extract more information from
it. For instance, we can add transparency (alpha) to avoid
overplotting:
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(alpha = 0.1)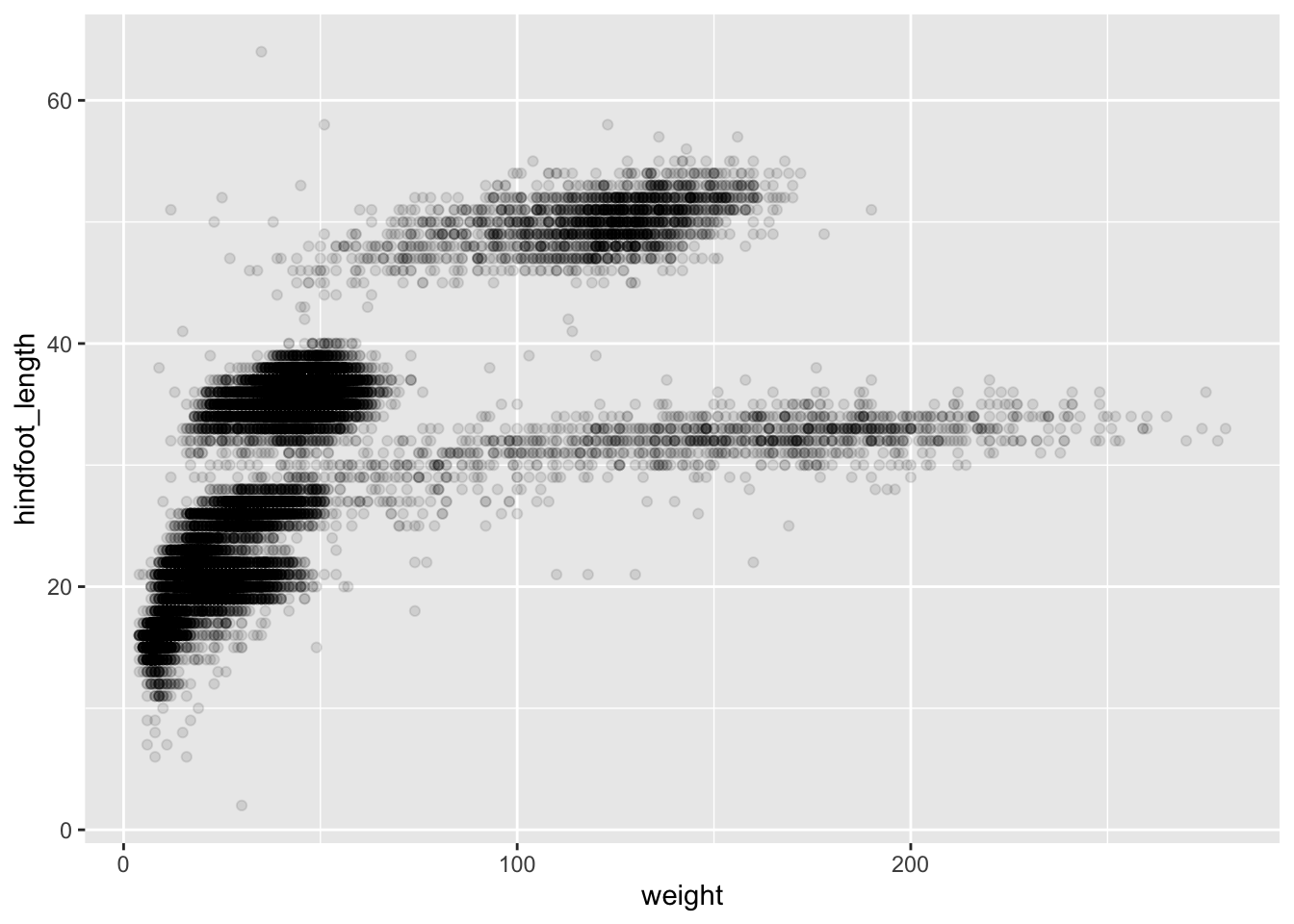
We can also add colors for all the points:
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(alpha = 0.1, color = "blue")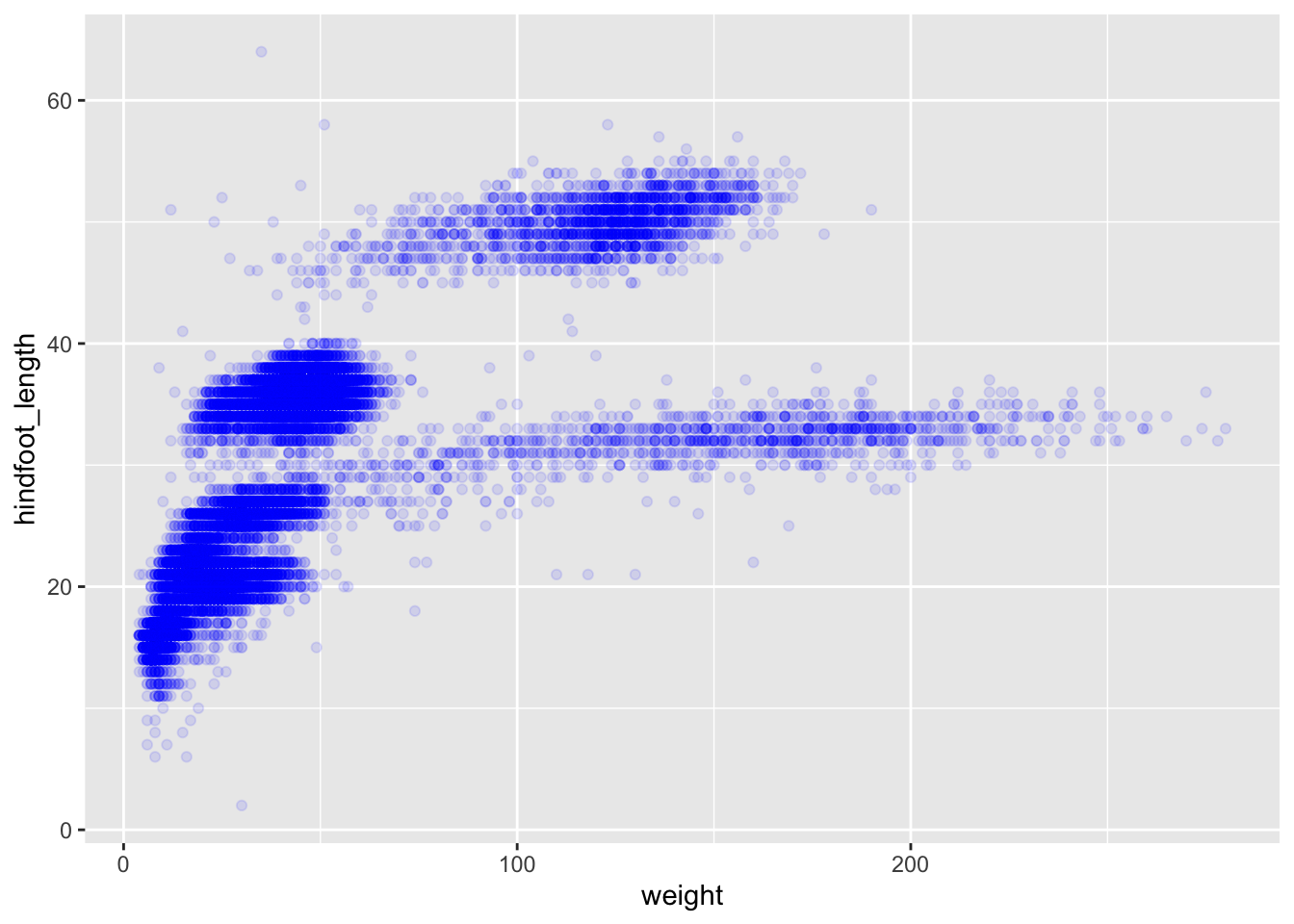
Or to color each species in the plot differently, you could use a
vector as an input to the argument color.
ggplot2 will provide a different color
corresponding to different values in the vector. Here is an example
where we color with species_id:
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length)) +
geom_point(alpha = 0.1, aes(color = species_id))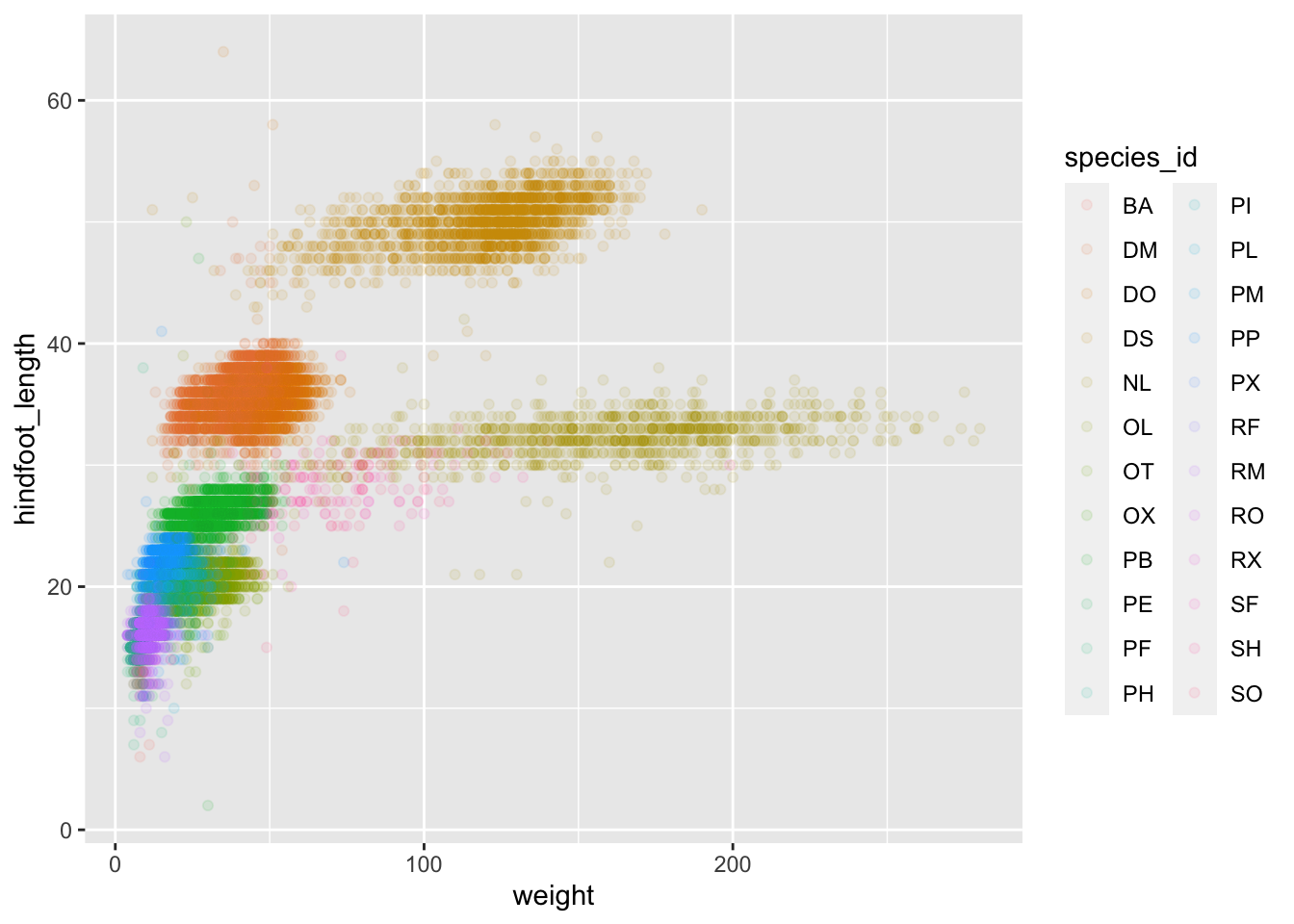
We can also specify the colors directly inside the mapping provided
in the ggplot() function. This will be seen by all geom
layers and the mapping will be determined by the x- and y-axis set up in
aes().
ggplot(data = surveys_complete, mapping = aes(x = weight, y = hindfoot_length, color = species_id)) +
geom_point(alpha = 0.1)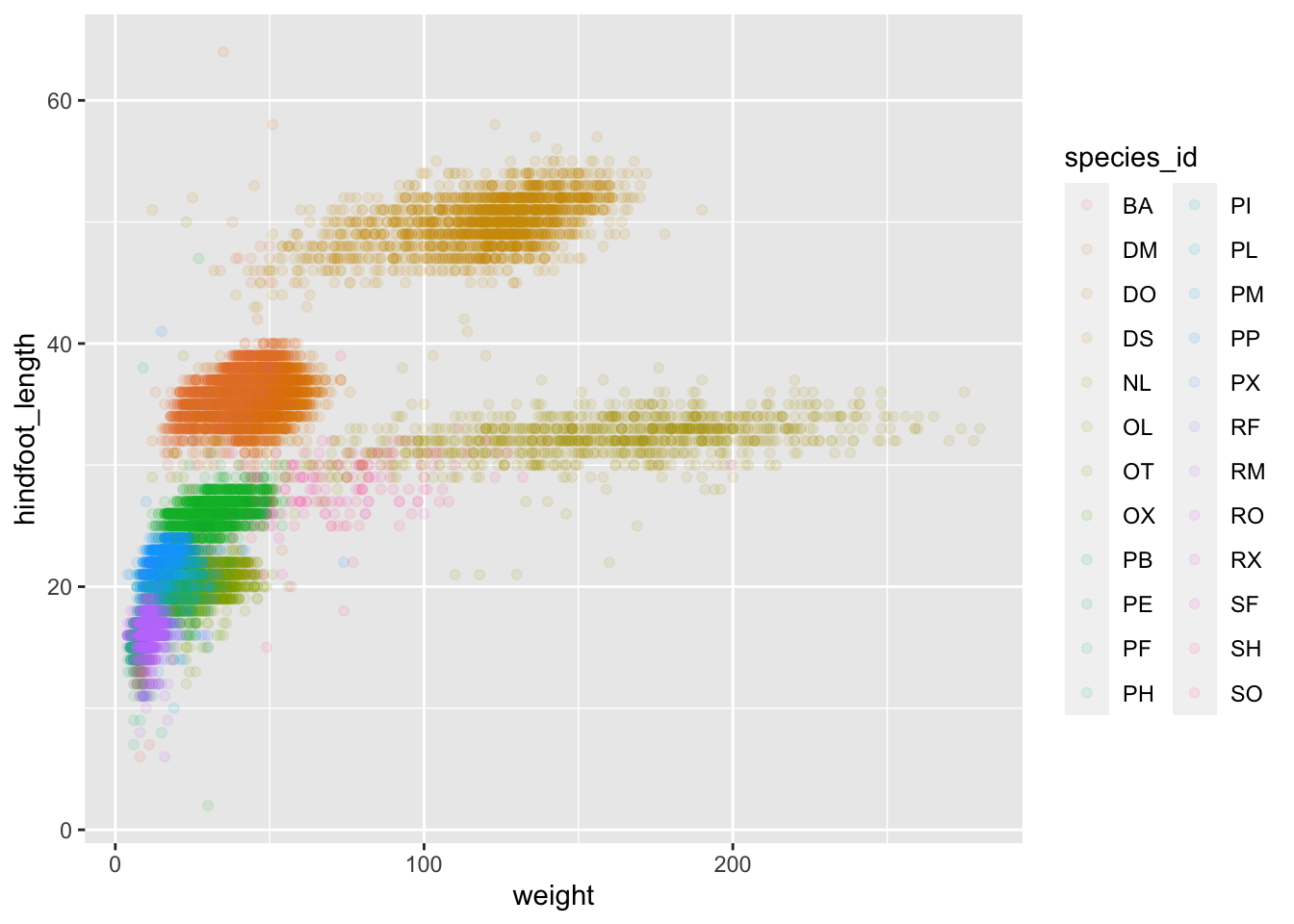
Notice that we can change the geom layer and colors will be still
determined by species_id
Challenge
Use ggplot() to create a scatter plot of
weight and species_id with weight on the
Y-axis, and species_id on the X-axis. Have the colors be coded by
plot_type. Is this a good way to show this type of data?
What might be a better graph?
ANSWER
ggplot(data = surveys_complete, mapping = aes(x = species_id, y = weight)) +
geom_point(aes(color = plot_type))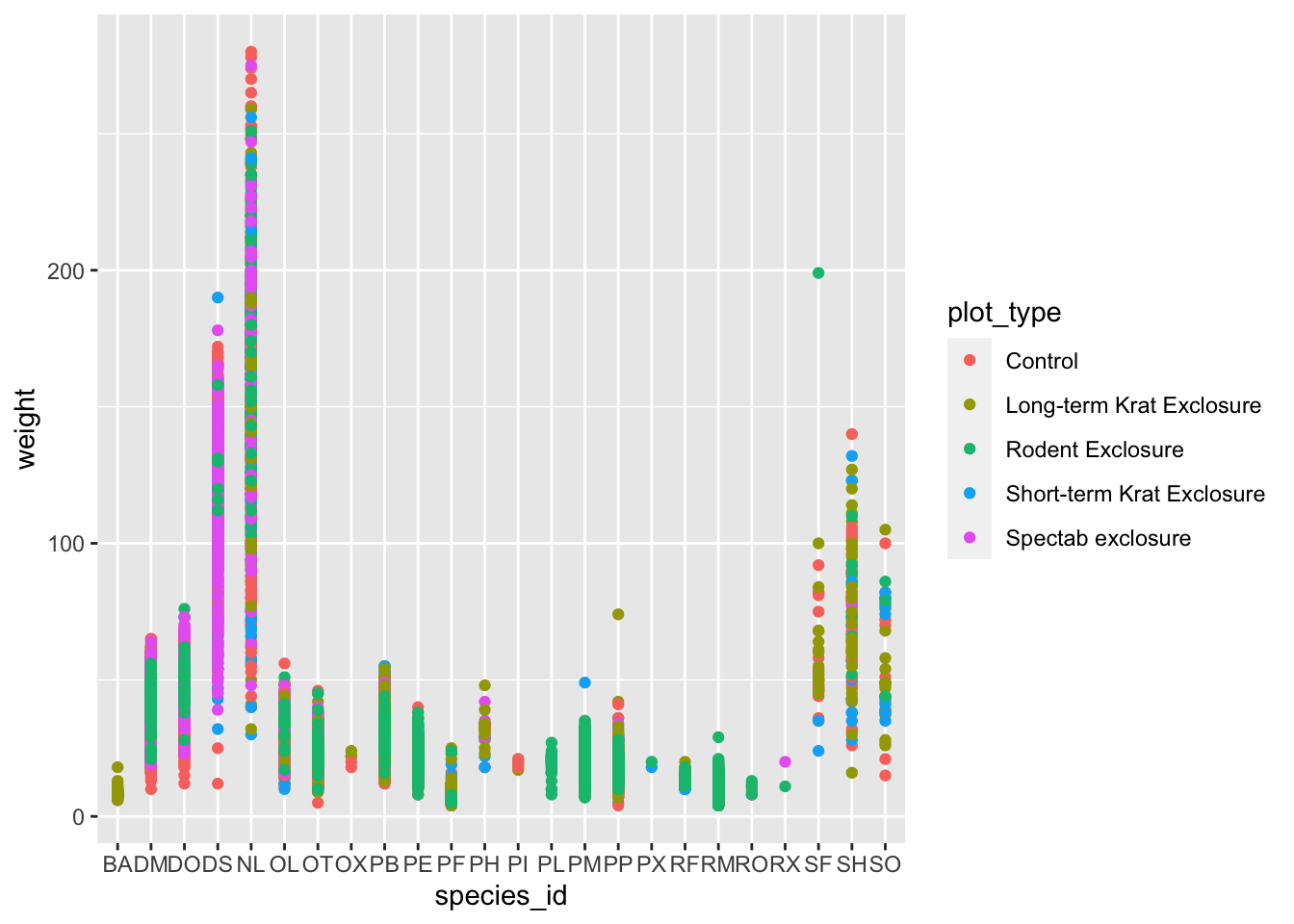
Boxplot
We can use boxplots to visualize the distribution of weight within each species:
ggplot(data = surveys_complete, mapping = aes(x = species_id, y = weight)) +
geom_boxplot()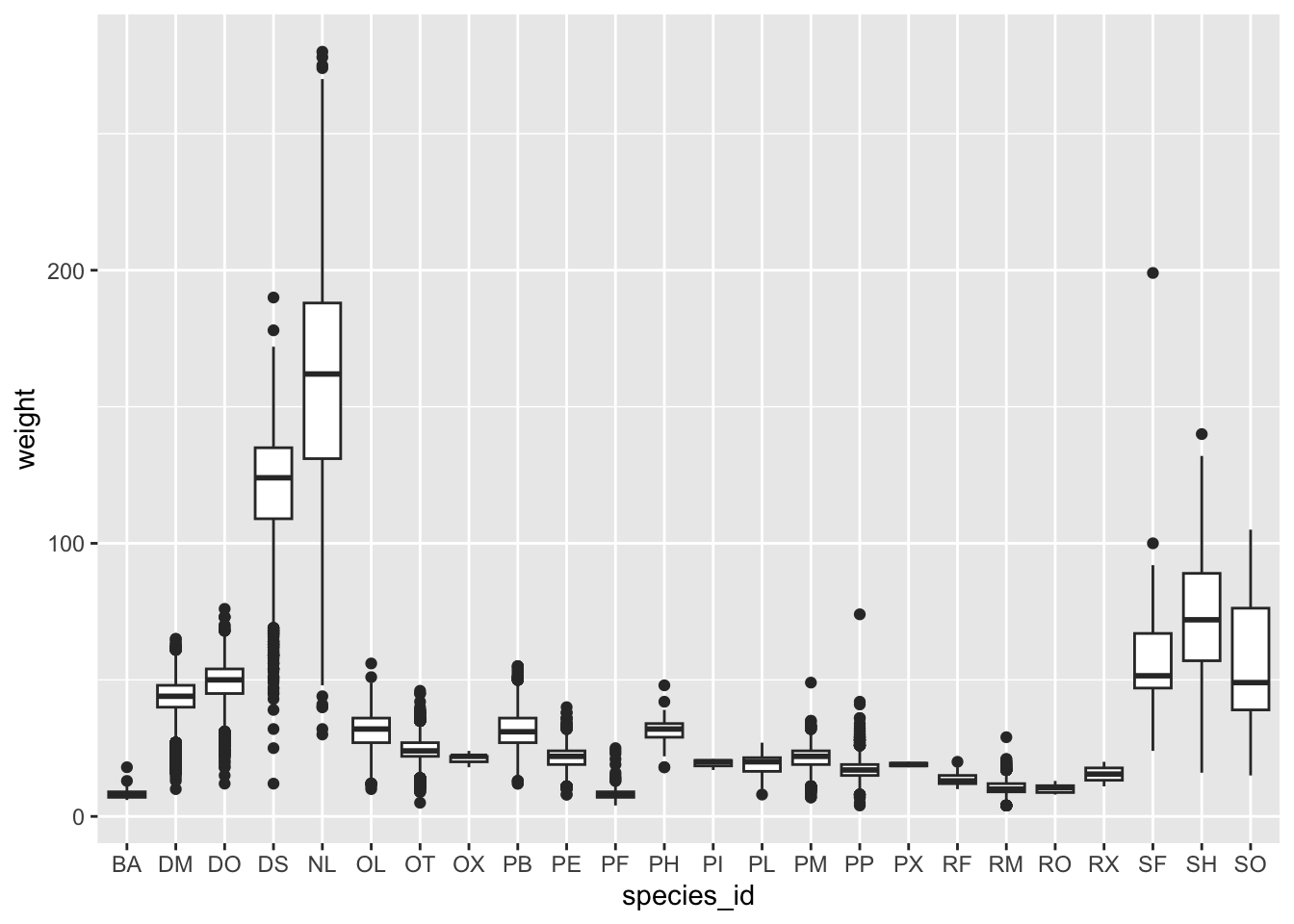
By adding points to boxplot, we can have a better idea of the number of measurements and of their distribution.
Let’s also use the geometry “jitter”. geom_jitter is
almost like geom_point but it allows you to visualize how
the density of points because it adds a small amount of random variation
to the location of each point.
ggplot(data = surveys_complete, mapping = aes(x = species_id, y = weight)) +
geom_boxplot(alpha = 0) +
geom_jitter(alpha = 0.3, color = "tomato") #notice our color needs to be in quotations 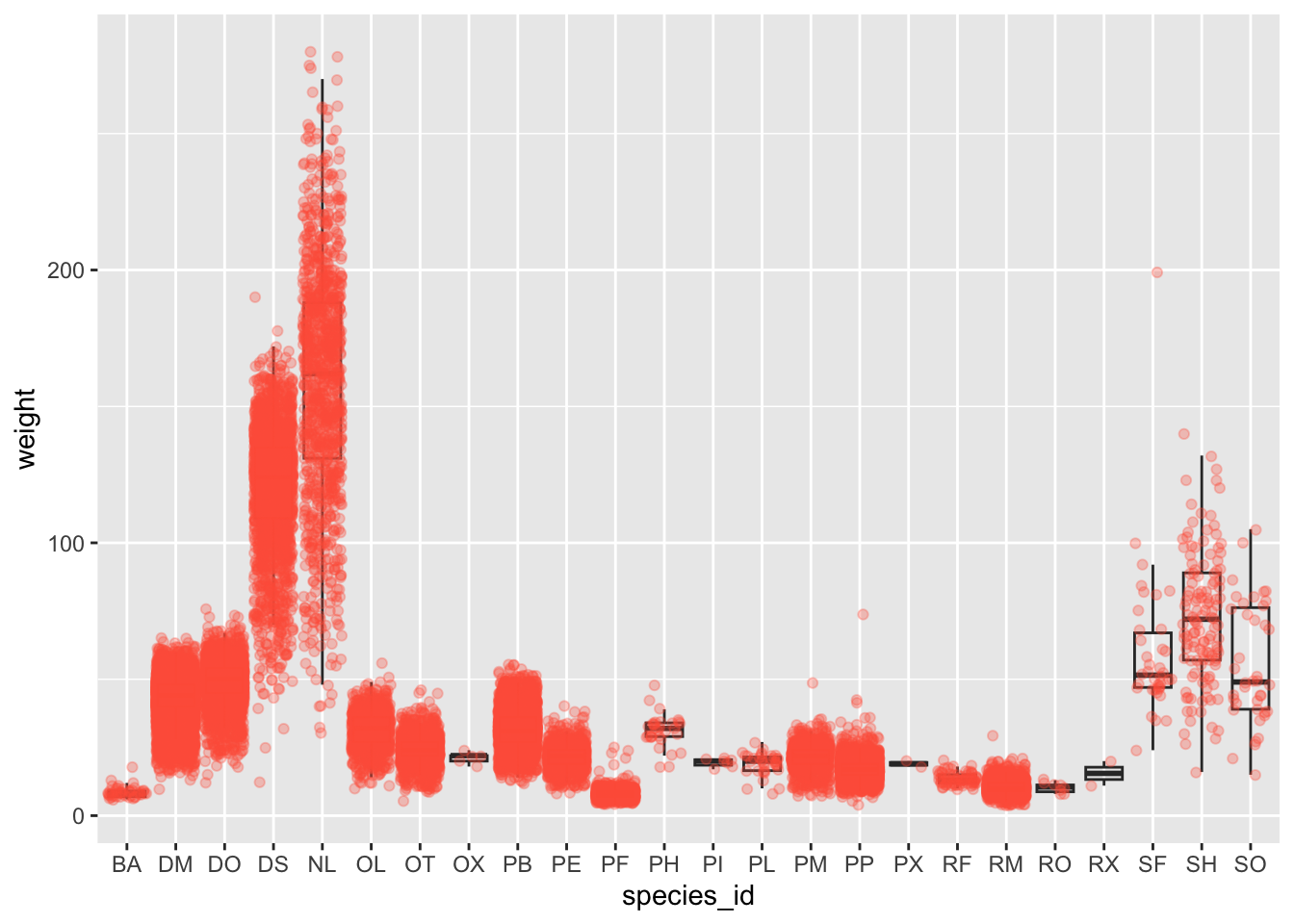
Notice how the boxplot layer is behind the jitter layer? What do you need to change in the code to put the boxplot in front of the points such that it’s not hidden?
Challenges
- Boxplots are useful summaries, but hide the shape of the distribution. For example, if the distribution is bimodal, we would not see it in a boxplot. An alternative to the boxplot is the violin plot, where the shape (of the density of points) is drawn.
- Replace the box plot code from above with a violin plot; see
geom_violin().
- In many types of data, it is important to consider the scale of the observations. For example, it may be worth changing the scale of the axis to better distribute the observations in the space of the plot. Changing the scale of the axes is done similarly to adding/modifying other components (i.e., by incrementally adding commands). Try making these modifications:
- Use the violin plot you made in Q1 and adjust the weight to be on
the log10 scale; see
scale_y_log10().
- Make a new plot to explore the distribution of
hindfoot_lengthjust for species NL and PF usinggeom_boxplot(). Overlay a jitter/scatter plot of the hindfoot lengths of the two species behind the boxplots. Then, add anaes()argument to color the datapoints (but not the boxplots) according to the plot from which the sample was taken.
Hint: Check the class for plot_id. Consider
changing the class of plot_id from integer to factor. Why
does this change how R makes the graph?
ANSWER
#1 + 2
ggplot(data = surveys_complete, mapping = aes(x = species_id, y = weight)) +
geom_jitter(alpha = 0.3, color = "tomato") +
geom_violin(alpha = 0) +
scale_y_log10()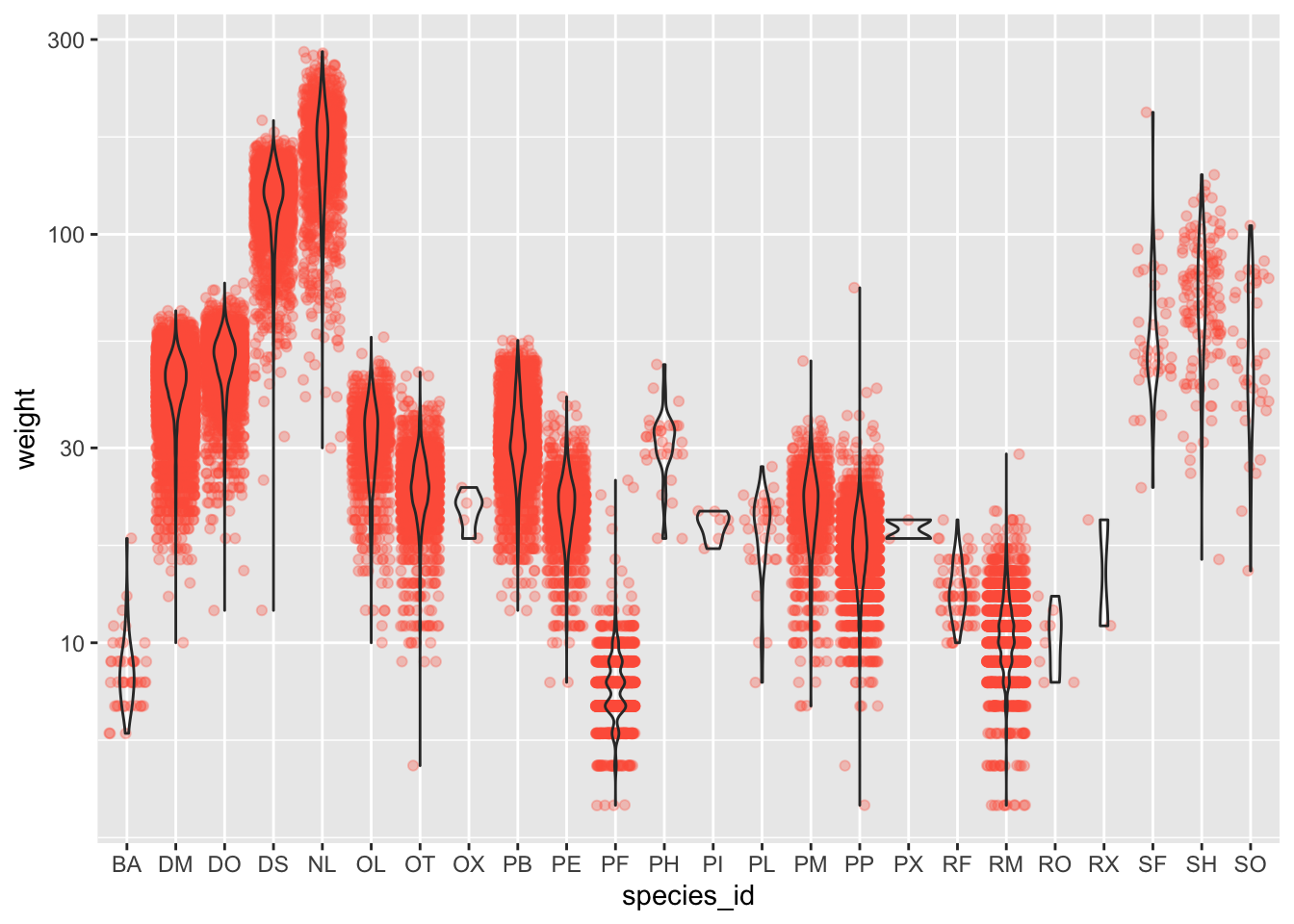
#3
surveys_complete %>%
filter(species_id == "NL" | species_id == "PF") %>%
ggplot(mapping = aes(x= species_id, y = hindfoot_length)) +
geom_jitter(alpha = 0.3, aes(color = as.factor(plot_id))) +
geom_boxplot()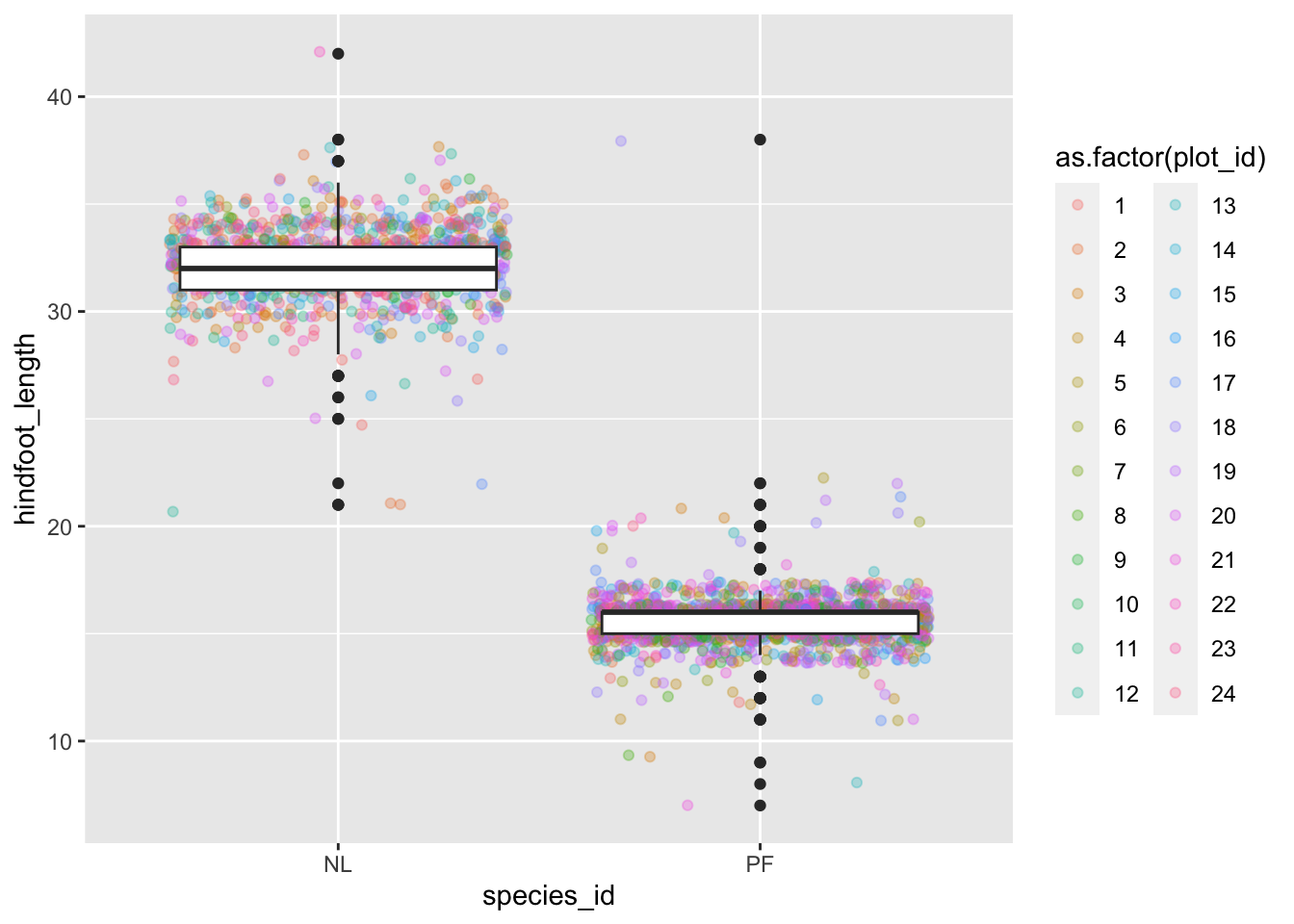
Plotting time series data
Let’s calculate number of counts per year for each species. First we
need to group the data and count records within each group. We can
quickly use the dplyr function count to do this.
count is very similar to the function tally we
have seen before, but it interally calls group_by before
the function and ungroup after.
yearly_counts <- surveys_complete %>%
count(year, species_id) Time series data can be visualized as a line plot with years on the x axis and counts on the y axis:
ggplot(data = yearly_counts, mapping = aes(x = year, y = n)) +
geom_line()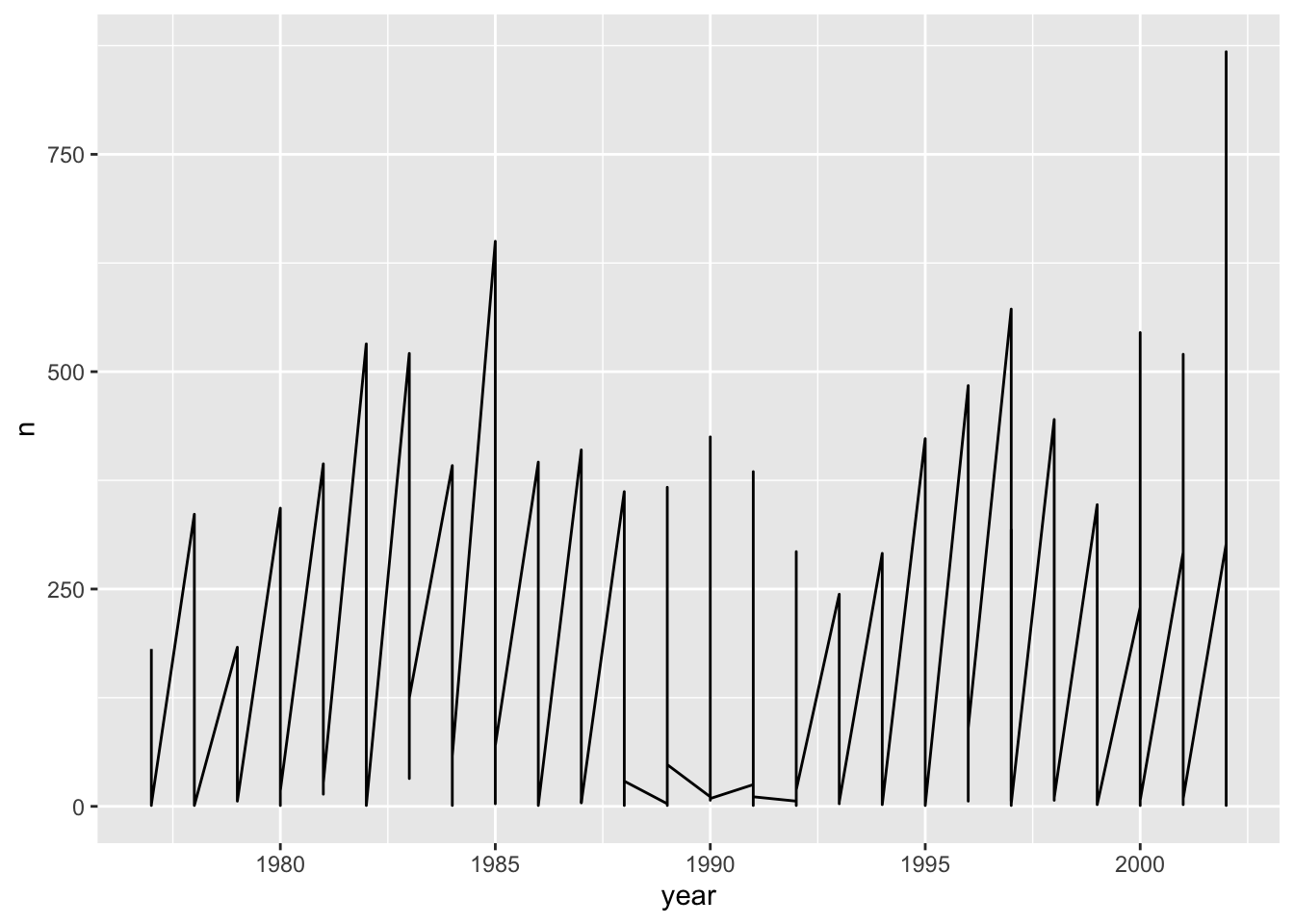
Unfortunately, this does not work because we plotted data for all the
species together. We need to tell ggplot to draw a line for each species
by modifying the aesthetic function to include
group = species_id:
ggplot(data = yearly_counts, mapping = aes(x = year, y = n, group = species_id)) +
geom_line()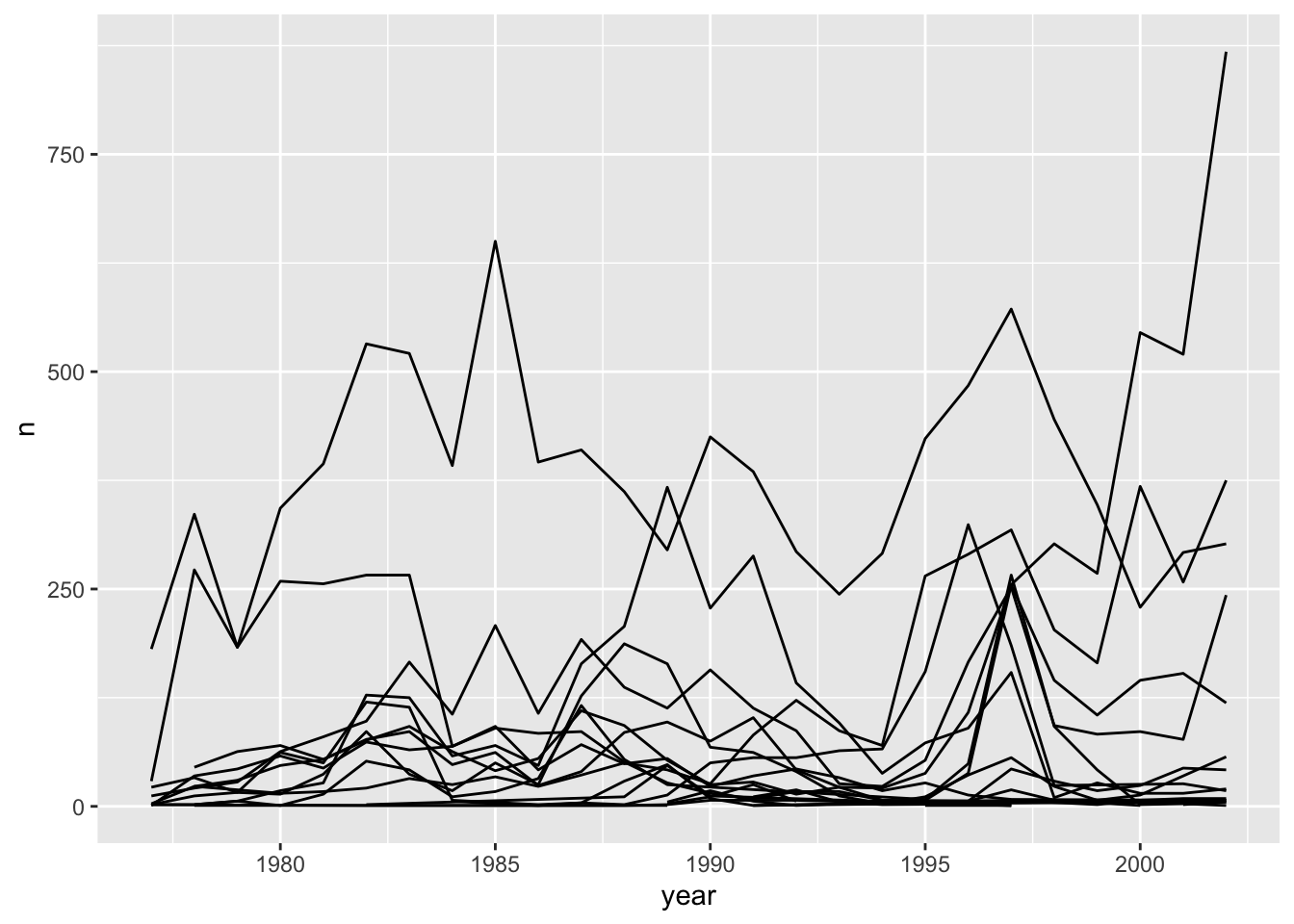
We will be able to distinguish species in the plot if we add colors
(using color also automatically groups the data):
ggplot(data = yearly_counts, mapping = aes(x = year, y = n, color = species_id)) +
geom_line()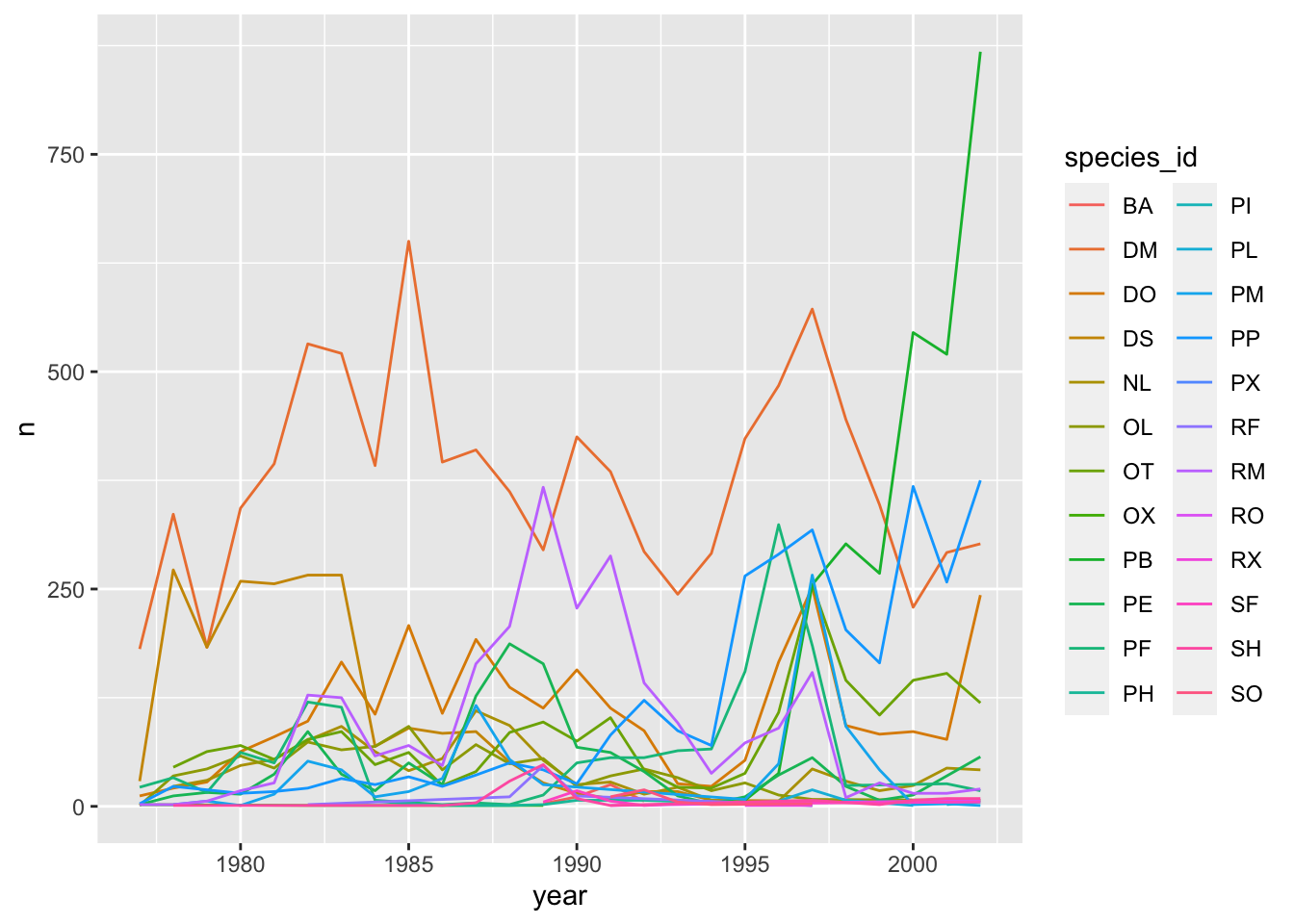
Faceting
ggplot2 has a special technique called
faceting that allows the user to split one plot into multiple
plots based on a factor included in the dataset. We will use it to make
a time series plot for each species:
ggplot(data = yearly_counts, mapping = aes(x = year, y = n)) +
geom_line() +
facet_wrap(~ species_id)## `geom_line()`: Each group
## consists of only one
## observation.
## ℹ Do you need to adjust the
## group aesthetic?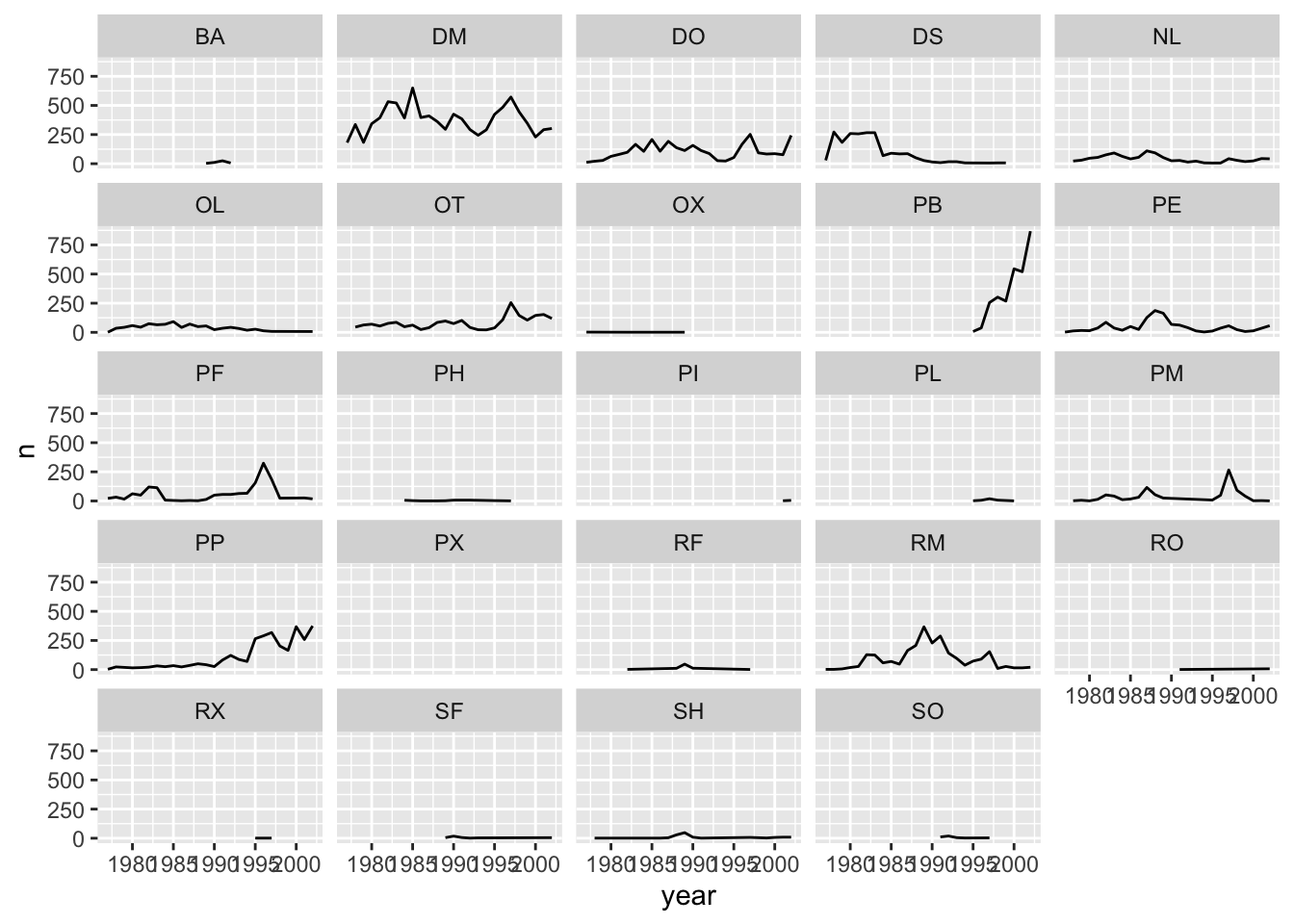
Challenge
You are looking at a new dataset shared with you by a collaborator. You received the dataset shortly after the vernal equinox. Your collaborator didn’t really give you any context on what the data represent, and you need to do some preliminary visualizations before you can really even formulate a question for them. Import the mystery dataset using:
mystery <- read_csv("https://raw.githubusercontent.com/gge-ucd/R-DAVIS/master/data/mysteryData.csv")## Rows: 91325 Columns: 3
## ── Column specification ──────
## Delimiter: ","
## chr (1): Group
## dbl (2): x, y
##
## ℹ Use `spec()` to retrieve the full column specification for this data.
## ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.Can you figure out what this dataset represents?
Hint Use your new knowledge of faceting to break up the data
into groups, and consider changing the size and transparency of your
geom_’s to get a better look!
ANSWER
# Preview the data
mystery %>%
head(5)## # A tibble: 5 × 3
## x y Group
## <dbl> <dbl> <chr>
## 1 414. 282. A
## 2 412. 281. A
## 3 414. 281. A
## 4 26.2 275. A
## 5 33.9 275. A# Plot the data
ggplot(data = mystery, mapping = aes(x = x, y = y)) +
facet_wrap(~ Group) +
geom_point(size = 0.1, alpha = 0.01) 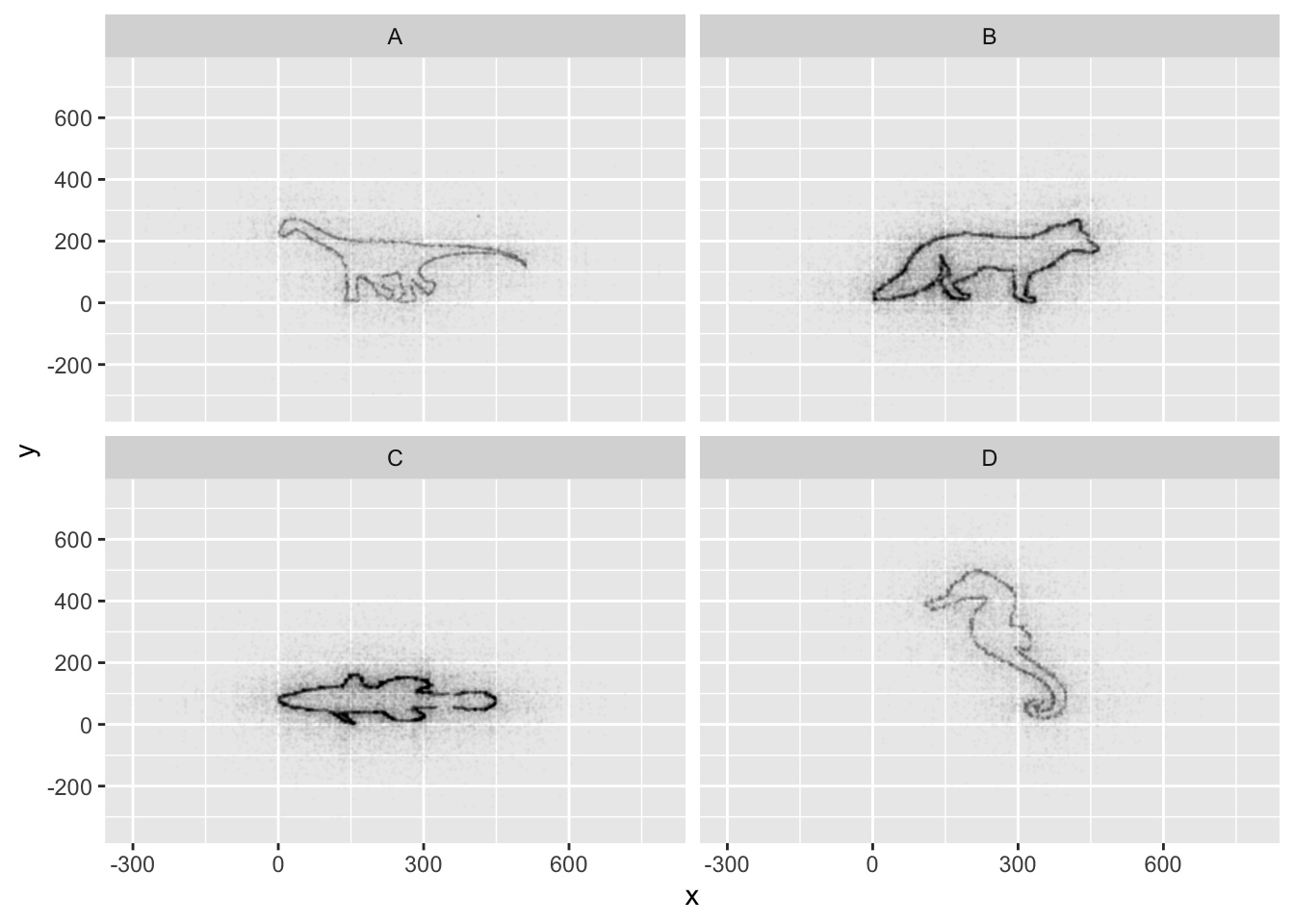
If all went well, and you faceted by group, set points to be very
small, and transparency to be very high (i.e., a low alpha
setting), you should discover that each group is the outline of a
different animal! You may notice that there is some distortion going on,
and our foxy friend in Group B appears to have some thick thighs. Try
equalizing the coordinate space in the x- and y-axes by adding a
coord_equal() to your ggplot() call:
## Equalize coordinate mapping
ggplot(data = mystery, mapping = aes(x = x, y = y)) +
facet_wrap(~ Group) +
geom_point(size = 0.1, alpha = 0.01) +
coord_equal()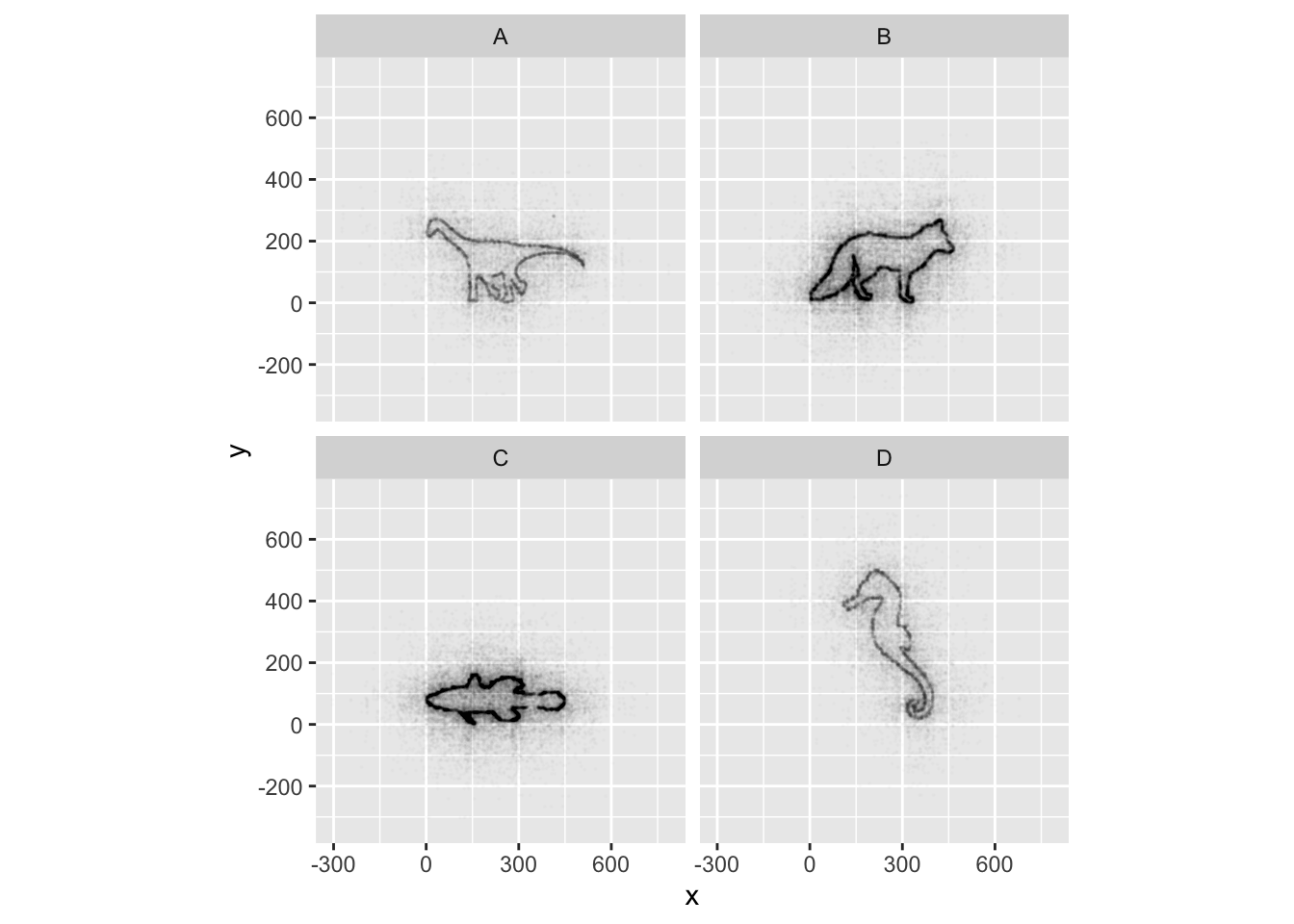
This challenge was inspired by Trevor Branch’s Coelocanth post on Twitter and adapted by Christian John.
ggplot2 themes
ggplot Themes are a great, easy addition that can make all your plots more readable (and a lot more pretty!)
In addition to theme_bw(), which changes the plot
background to white, ggplot2 comes with
several other themes which can be useful to quickly change the look of
your visualization. The complete list of themes is available at http://docs.ggplot2.org/current/ggtheme.html.
theme_minimal() and theme_light() are popular,
and theme_void() can be useful as a starting point to
create a new hand-crafted theme.
Usually plots with white background look more readable when printed.
We can set the background to white using the function
theme_bw(). Additionally, you can remove the grid:
ggplot(data = yearly_counts, mapping = aes(x = year, y = n)) +
geom_line() +
facet_wrap(~ species_id) +
theme_bw() +
theme(panel.grid = element_blank())## `geom_line()`: Each group
## consists of only one
## observation.
## ℹ Do you need to adjust the
## group aesthetic?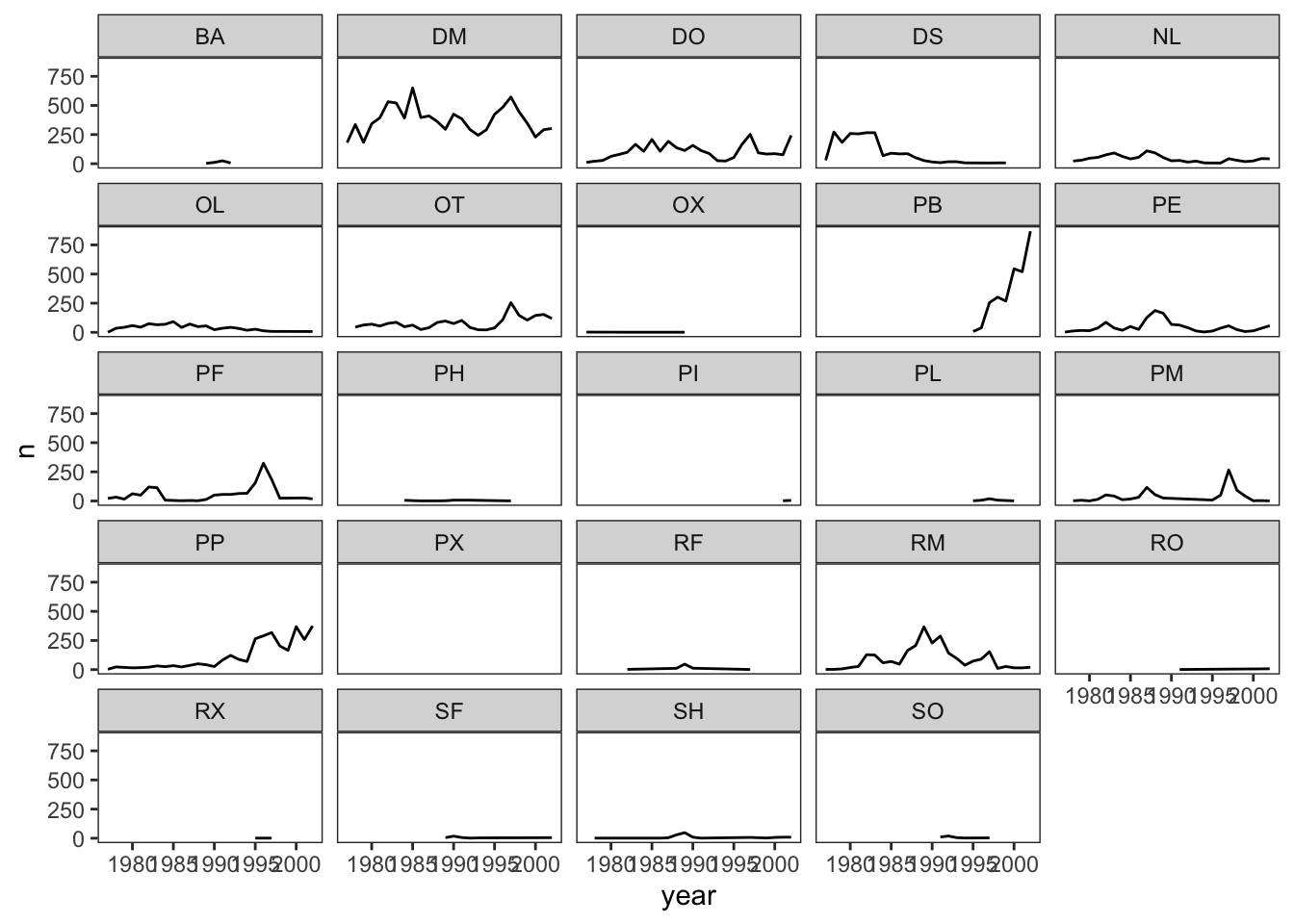
Challenge 1
Let’s make one final change to our facet wrapped plot of our yearly count data. What if we wanted to split the counts of species up by sex where the lines for each sex are different colors? Make sure you have a nice theme on your graph too!
Hint Make a new dataframe using the count
function we learned earlier!
ANSWER
#new data frame counting the number of each sex of each species
yearly_sex_counts <- surveys_complete %>%
count(year, species_id, sex)
#plot code
ggplot(data = yearly_sex_counts, mapping = aes(x = year, y = n, color = sex)) +
geom_line() +
facet_wrap(~ species_id) +
theme_bw()## `geom_line()`: Each group
## consists of only one
## observation.
## ℹ Do you need to adjust the
## group aesthetic?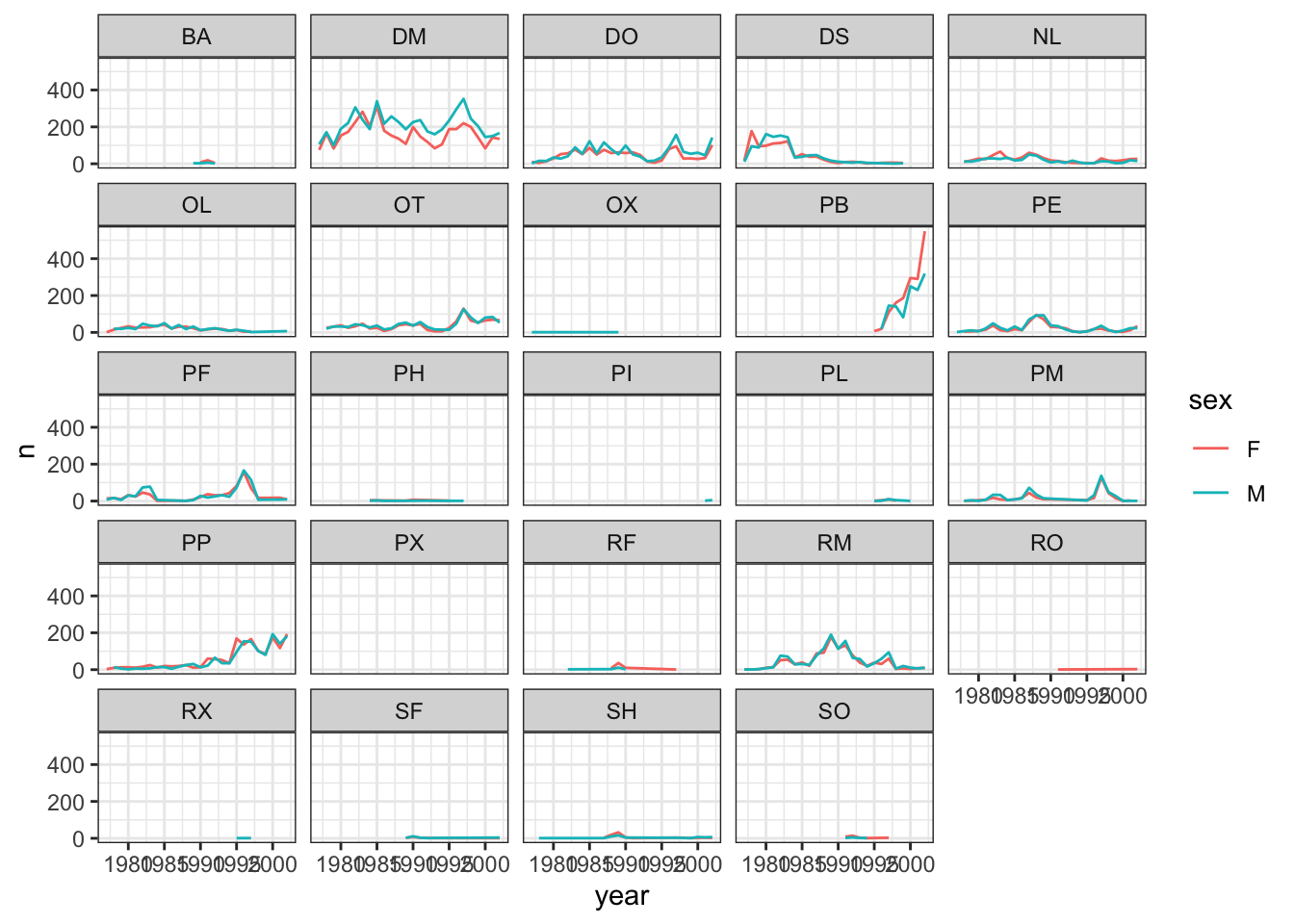
Challenge 2
Use what you just learned to create a plot that depicts how the average weight of each species changes through the years.
ANSWER
#create a new dataframe
yearly_weight <- surveys_complete %>%
group_by(year, species_id) %>%
summarize(avg_weight = mean(weight))## `summarise()` has grouped
## output by 'year'. You can
## override using the `.groups`
## argument.ggplot(data = yearly_weight, mapping = aes(x=year, y=avg_weight)) +
geom_line() +
facet_wrap(~ species_id) +
theme_bw()## `geom_line()`: Each group
## consists of only one
## observation.
## ℹ Do you need to adjust the
## group aesthetic?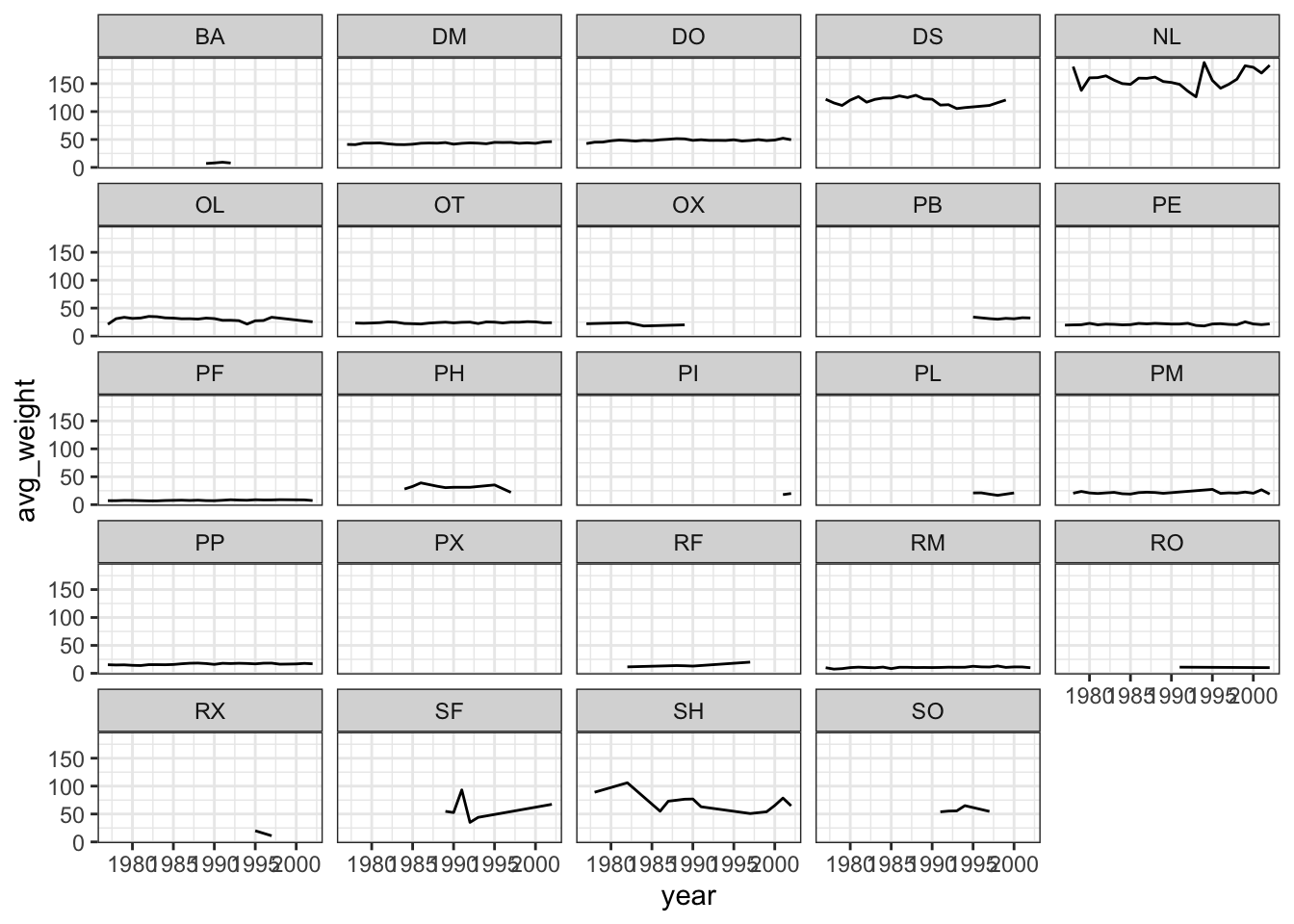
This lesson was contributed by Martha Zillig.