Creating functions
Learning objectives
- Define a function that takes arguments.
- Set default values for function arguments.
- Explain why we should divide programs into small, single-purpose functions.
Any operation you will perform more than once can be put into a
function. That way, rather than retyping all the commands (and
potentially making errors), you can simply call the function, passing it
a new dataset or parameters. This may seem cumbersome at first, but
writing functions to automate repetitive tasks is incredibly powerful.
E.g. each time you call ggplot you are calling a function
that someone wrote. Imagine if each time you wanted to make a plot you
had to copy and paste or write that code from scratch!
Defining a function
Recall the components of a function. E.g. the log
function (see ?log) takes “arguments” x and
base and “returns” the base-base logarithm of
x. Functions take arguments as input and yield
return-values as output. You can define functions to do any number of
operations on any number of arguments, but always output a single return
value (however there are complex objects into which you can put multiple
objects, should you need to).
Let’s start by defining a simple function to add two numbers. This is
the basic structure, which you can read as “assign to the variable
my_sum a function that takes arguments a and
b and returns the_sum.” The body of the
function is delimited by the curly-braces. The statements in the body
are indented. This makes the code easier to read but does not affect how
the code operates.
my_sum <- function(a, b) {
the_sum <- a + b
return(the_sum)
}Notice that no numbers were summed when we ran that code, but now the
Environment has an object called my_sum that has type
function. You can call my_sum just like you would any other
function. When you do, the code between the curly-braces of the
my_sum definition is run with whatever values you pass to
a and b substituted in their place.
my_sum(a = 2, b = 2)## [1] 4my_sum(3, 4)## [1] 7Just like log provides a default value of
base (exp(1)) so that you don’t have to type
it every time, you can provide default values to any arguments of your
function. Then if the user doesn’t specify them, the defaults will be
used.
my_sum2 <- function(a = 1, b = 2) {
the_sum <- a + b
return(the_sum)
}
my_sum2()## [1] 3my_sum2(b = 7)## [1] 8Tip
One feature unique to R is that the return statement is not required.
R automatically returns the output of the last line of the body of the
function unless a return statement is specified elsewhere.
Since other languages require a return statement and
because it can make reading a function easier, we will explicitly define
the return statement.
Temperature conversion
Let’s define a function F_to_K that converts temperatures from Fahrenheit to Kelvin:
F_to_K <- function(temp) {
K <- ((temp - 32) * (5 / 9)) + 273.15
return(K)
}Calling our own function is no different from calling any other function:
# freezing point of water
F_to_K(32)## [1] 273.15# boiling point of water
F_to_K(212)## [1] 373.15Challenge
- Write a function called
K_to_Cthat takes a temperature in K and returns that temperature in C- Hint: To convert from K to C you subtract 273.15
- Create a new R script, copy
F_to_KandK_to_Cin it, and save it as functions.R in thecodedirectory of your project.
ANSWER
K_to_C <- function(tempK) {
tempC <- tempK - 273.15
return(tempC)
}source()ing functions
You can load all the functions in your code/functions.R
script without even opening the file, via the source
function. This allows you to keep your functions separate from the
analyses which use them.
source('code/functions.R')Using dataframes in functions
Let’s write a function to calculate the average GDP in a given
country, in a given span of years, based on the gapminder
data. If you were to do this for just one specification, without writing
a function, it might look something like this:
library(gapminder)
library(tidyverse)
gapminder %>%
filter(country == "Canada", year %in% c(1950:1970)) %>%
summarize(mean(gdpPercap, na.rm = T))## # A tibble: 1 × 1
## `mean(gdpPercap, na.rm = T)`
## <dbl>
## 1 13349.But, what if you wanted to do this for many different specifications? You might find yourself wanting to copy and paste these couple lines of code over and over. Instead, you can write it into a function, soft coding the parts that you want as your function’s arguments. In this example, we want to be able to change the country and the year range that we are interested in.
# Note: try to name arguments something that do not exist as a column name, to avoid confusing yourself and R
avgGDP <- function(cntry, yr.range){
df <- gapminder %>%
filter(country == cntry, year %in% yr.range)
mean(df$gdpPercap, na.rm = T)
}
avgGDP("Iran", 1980:1985)## [1] 7608.335avgGDP("Zimbabwe", 1950:2000)## [1] 648.8549Pass by value
Functions in R almost always make copies of the data to operate on inside of a function body. When we modify a data frame inside the function we are modifying the copy of the gapminder dataset, not the original variable we gave as the first argument. This is called “pass-by-value” and it makes writing code much safer: you can always be sure that whatever changes you make within the body of the function, stay inside the body of the function.
Challenge
This challenge will deal with countries’ population growth. To access
the data, load (and install, if needed) the gapminder
library and access its life expectancy dataset using:
library(gapminder)
d <- gapminder::gapminderWrite a new function that takes two arguments, the gapminder
data.frame (d) and the name of a country
(e.g. "Afghanistan"), and plots a time series of the
country’s population. The return value from the function should be a
ggplot object. Note: It is often easier to modify existing code than to
start from scratch. To start out with one plot for a particular country,
figured out what you need to change for each iteration (these will be
your arguments), and then wrap it in a function.
ANSWER
plotPopGrowth <- function(countrytoplot, dat = gapminder) {
df <- filter(dat, country == countrytoplot)
plot <- ggplot(df, aes(year, pop)) +
geom_line()
return(plot)
}
plotPopGrowth('Canada')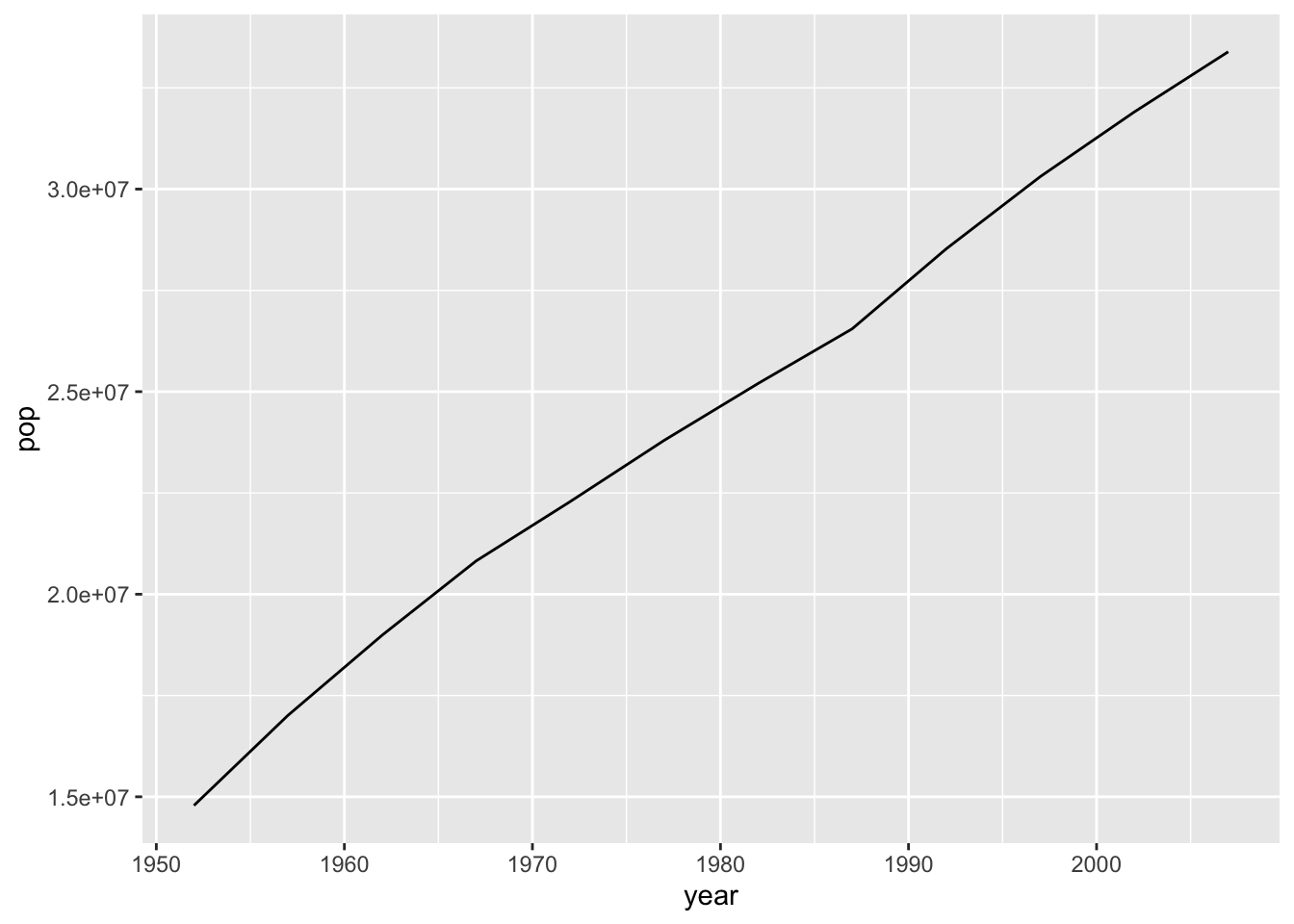
This lesson is adapted from the Software Carpentry: R for Reproducible Scientific Analysis Creating Functions materials.